Geom_segment(xend = 0, yend = 0, arrow = arrow_style) +Ĭoord_fixed() + # fix aspect ratio to 1:1 Pivot_wider(names_from = "PC", names_prefix = "PC", values_from = "value") %>% Now in the context of a plot: # define arrow style for plotting Tidy(matrix = "rotation") # A tibble: 81 x 3 When applied to prcomp objects, the tidy() function takes an additional argument matrix, which we set to matrix = "rotation" to extract the rotation matrix. The rotation matrix is stored as pca_fit$rotation, but here we’ll extract it using the tidy() function from broom. pca_fit %>%Īugment(biopsy) %>% # add original dataset back in The columns containing the fitted coordinates are called. We do this with the augment() function from broom, which takes as arguments the fitted model and the original data.
#Add pca column back to data Pc#
In general, this means combining the PC coordinates with the original dataset, so we can color points by categorical variables present in the original data but removed for the PCA. Now, we want to plot the data in PC coordinates.
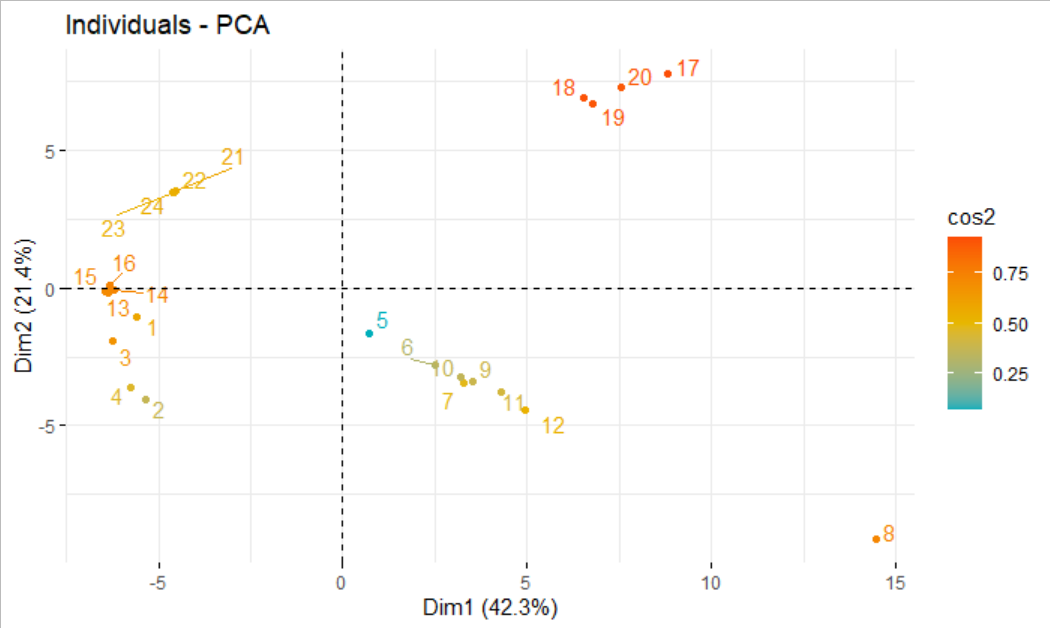
Prcomp(scale = TRUE) # do PCA on scaled dataĪs an alternative to scale = TRUE, we could also have scaled the data by explicitly invoking the scale() function. Select(where(is.numeric)) %>% # retain only numeric columns We do so by using the argument scale = TRUE in prcomp(). Second, we normally want to scale the data values to unit variance before PCA.
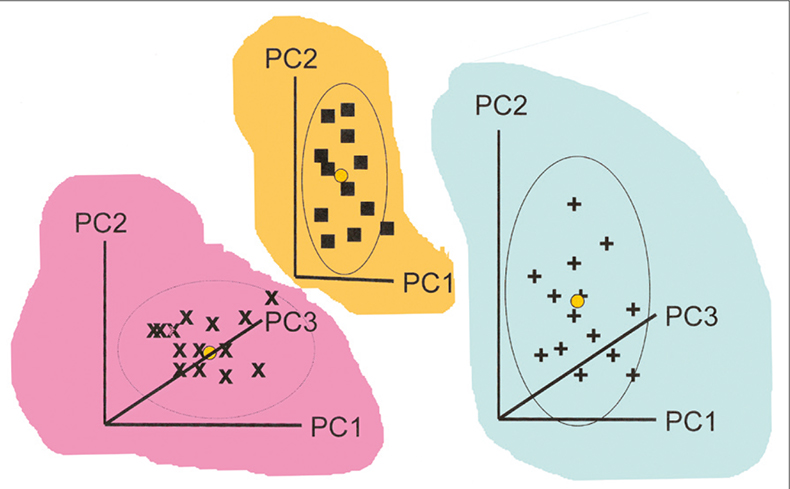
This is straightforward using the where(is.numeric) tidyselect construct. First, the prcomp() function can only deal with numeric columns, so we need to remove all non-numeric columns from the data. We start by running the PCA and storing the result in a variable pca_fit. Look at the variance explained by each PC.In general, when performing PCA, we’ll want to do (at least) three things:
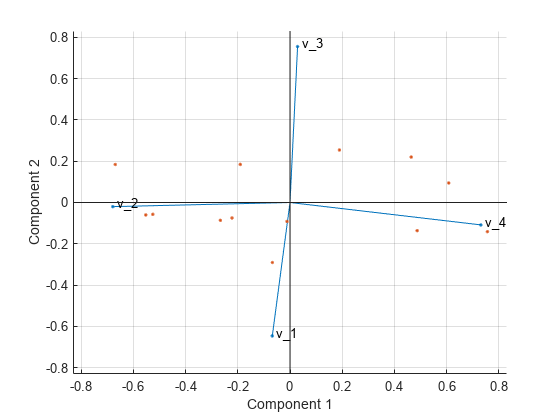
biopsy <- read_csv("") # Parsed with column specification: The true outcome (benign/malignant) is also known. He assessed biopsies of breast tumors for 699 patients each of nine attributes was scored on a scale of 1 to 10. It is a breast cancer dataset from the University of Wisconsin Hospitals, Madison from Dr. William H. We’ll be analyzing the biopsy dataset, which comes originally from the MASS package. # x dplyr::lag() masks stats::lag() library(broom) # devtools::install_github("tidymodels/broom") # x dplyr::filter() masks stats::filter() We’ll also use the cowplot package for plot themes. Note that as of this writing, we need the current development version of broom because of a bug in tidy.prcomp(). But now, I’ve realized that all the necessary functions to do this tidyverse-style are available in the broom package.įor our PCA example, we’ll need the packages tidyverse and broom. While it’s reasonably easy to extract the relevant info with some base-R manipulations, I’ve never been happy with this approach. The result is an object of class prcomp that doesn’t fit nicely into the tidyverse framework, e.g. for visualization.
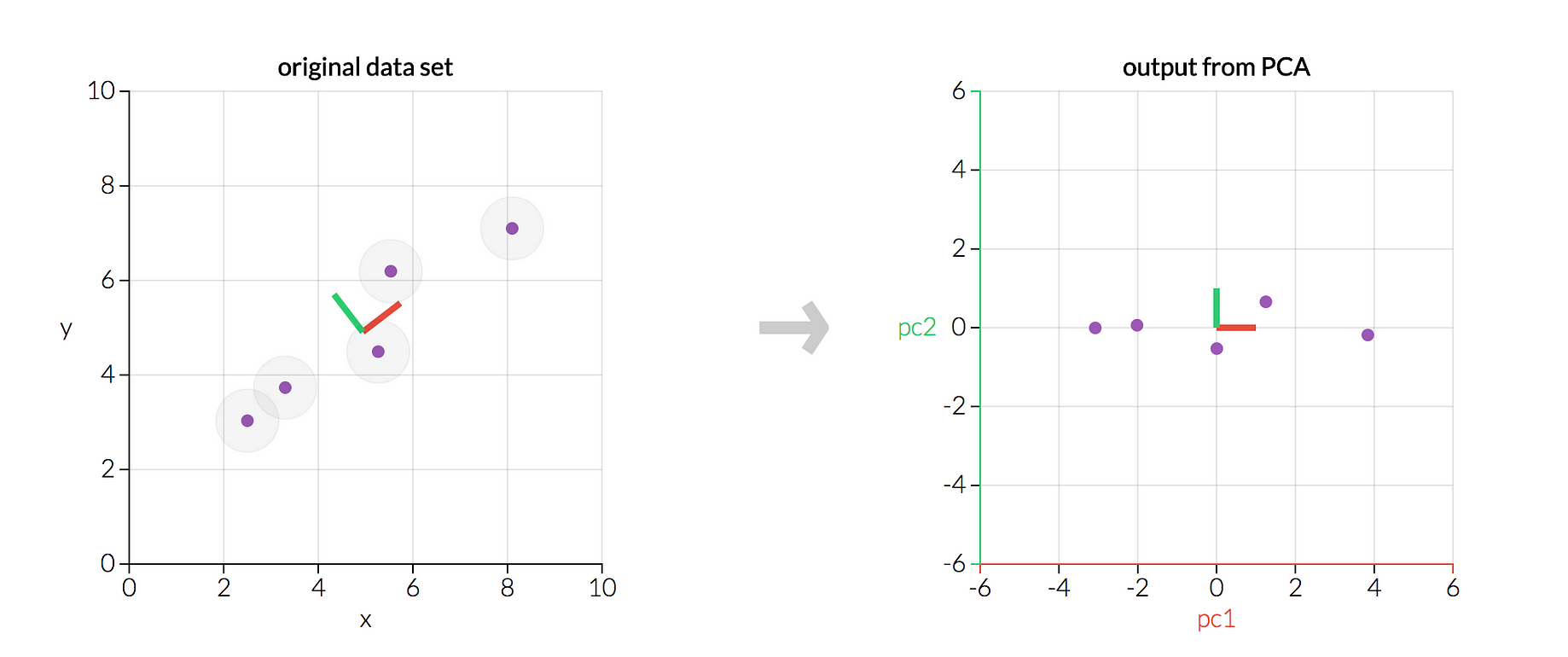
Doing a PCA in R is easy: Just run the function prcomp() on your matrix of scaled numeric predictor variables.
